Ultima Dual-mode mRNA Library Prep Kit for Illumina
Product Description
Hieff NGS™ Ultima Dual-mode mRNA Library Prep Kit for Illumina contains two independent modules. The core components of BOX-I are oligo (dT) magnetic beads required for mRNA purification. BOX-II contains mRNA fragmentation reagents, reverse transcription reagents, conventional and chain-specific ds-cDNA synthesis reagents, and others required for subsequent library construction. Therein dTTP is replaced with dUTP in the strand-specific two-strand synthesis Buffer, so that dUTP is incorporated into the second strand of cDNA. While the high-fidelity DNA polymerase used in this kit cannot amplify the DNA template containing uracil, achieving strand specificity. All reagents provided have undergone strict quality control and functional verification, ensuring the stability and reproducibility of library construction to the greatest extent.
Shipping and Storage
The Hieff NGS™ Ultima Dual-mode mRNA Library Prep Kit for Illumina® components in Box I are shipped with ice packs and can be stored at 2-8°C for one year.
The Hieff NGS™ Ultima Dual-mode mRNA Library Prep Kit for Illumina® components in Box II are shipped with dry ice and can be stored at -20°Cfor one year.
Cautions
1 Operation
1.1 For your safety and health, please wear personal protective equipment (PPE), such as laboratory coats and disposable gloves, when operating with this product. This product is for research use ONLY!
1.2 Thaw components at room temperature. Mix thoroughly by inverting up and down several times, spin down briefly and place on ice for use.
1.3 It is recommended to perform each step reaction in a thermal cycler with a heated lid. The thermal cycler should be preheated to the set temperature prior to use.
1.4 Supplies free of RNase contamination and cleaning the experimental area regularly are necessary. ThermoFisher's RNAZapTM high-efficiency nucleic acid removal spray was recommended to remove RNase contamination.
1.5 Improper operations may very likely cause aerosol contaminations, impacting the accuracy of the result. Mandatory physical isolation of PCR reaction mixing regions and PCR product purification assay regions is recommended. Equipped with equipment such as specialized pipettes for library construction..
2 Application
2.1 For research use only!
2.2 This kit is suitable for high-quality total RNA from eukaryotes such as animals, plants, and fungi with a starting input of 10 ng-4 μg (volume≤50 μL). If the initial RNA concentration is low and the volume exceeds 50 μL, it is recommended to condensed the DNA with Hieff NGS™ RNA Cleaner magnetic beads. To ensure that the mRNA has a complete poly(A) tail structure, RNA needs to be detected by the Agilent 2100 Bioanalyzer RNA 6000 Nano/Pico chip and the RIN value must be > 7,
2.3 Oligo (dT) magnetic beads was applied in the mRNA isolation module of this kit So that only mRNA with poly(A) tail can be extracted; other RNAs without poly(A) tail, such as non-coding RNA, no poly(A) Tail mRNA etc. were washed away. In addition, this kit is not compatible with FFPE samples since the mRNA in the FFPE sample is severely degraded and usually does not have a complete poly(A) tail structure.
2.4 The library prepared by this kit can be applied to a variety of RNA-Seq, including:
- Gene expression
- Single nucleotide variation discovery
- Gene fusion identification
- Splice variant analysis
3 Adapter Ligation
3.1 Yeasen provides long adapter (Barcoded Adapter) kits and short adapter (also called small Y adapters, incomplete adapters) kits, whichcustomers can choose according to experimental needs. There are currently 48 Indexed Adapters: Hieff NGS™ Complete Adapter Kit for Illumina®, Set 1~Set 4 (Cat#12615-Cat#12618); Double-ended Index Primers: Hieff NGS™ RNA 384 CDI Primer Kit for Illumina®, Set 1~Set 2 (Cat#12414-12415).
3.2 Selecting high-quality, commercial adapters was recommended. If self-made adapters are selected, please entrust a company with experience in NGS primer synthesis and remark on the need for strict contamination control. In addition, it is recommended to prepare DNA annealing solution in a clean bench and only operate one type of adapter each time to prevent cross-contamination.
3.3 Please thaw the adapters on the ice or at 4°C; when operating at room temperature, the laboratory temperature should not exceed 25°C to prevent the adapters from denaturing.
3.4 The concentration of the adapter directly affects the ligation efficiency and library yield. The adaptor volume added to the kit is fixed to 5ul. The adapters are recommended to be diluted with 0.1×TE buffer and the diluted adapters can be stored at 4°C for 48 hours. Table 1 lists the recommended adapter amount for different amounts of input RNA.
Table 1. The recommended adapter amount for different input RNA
|
Input Total RNA |
Adapter stock concentration |
|
10 ng |
1 μM |
|
100 ng |
1.5 μM |
|
500 ng |
3 μM |
|
≥1 μg |
5 μM |
4 Bead-based DNA Cleanup and Size Selection
4.1 There are multiple steps in the library construction process that require DNA purification magnetic beads. We recommend Hieff NGS™ DNA Selection Beads (Yeasen Cat#12601) or AMPure® XP magnetic beads (Beckman Cat#A63880) for DNA purification and size-selection.
4.2 The magnetic beads should be equilibrated at room temperature prior to use, otherwise the yield will decrease and the size-selecting effect will be affected.
4.3 The magnetic beads should be mixed well by vortex or pipetting prior to use.
4.4 Do not aspirate the beads when transferring the supernatant, even trace amounts of the beads may impact the following reactions.
4.5 The 80% ethanol should be freshly prepared, otherwise it will affect the recovery efficiency.
4.6 The magnetic beads should be dried at room temperature before the product is eluted. Insufficient drying will easily cause residual ethanol to affect subsequent reactions; excessive drying will cause the magnetic beads tocrack and reduce the purification yield. Normally, drying at room temperature for 3-5 minutes is enough to allow the beads to fully dry.
4.7 If needed, the purified or size-selected DNA samples eluted in TE buffer can be stored at 4°C for 1-2 weeks or at -20°C for a month.
5 Library Amplification
5.1 On the basis of the first-generation DNA polymerase, the high-fidelity DNA polymerase in the kit has greatly improved its amplification uniformity and exhibits no amplification bias.
5.2 If Indexed Adapter (also known as a long adapter or large Y adapter) is ligated to the target DNA, the primer mix provided in this kit can be used for amplification; if a "short adapter" or "small Y adapter" is used for DNA ligation, index primers are needed for amplification.
5.3 Amplification cycle numbers should be strictly controlled. Insufficient amplification may lead to low library yield; Over-amplification may introduce increased bias, errors, duplicated read, chimeric products, and accumulation of expansion mutations. Table 2 lists the recommended cycle numbers for PCR amplification.
Table 2 The recommended number of cycles to generate RNA library *
|
Input Total RNA |
Number of cycles |
|
|
Non-stranded |
Stranded |
|
|
10 ng |
15 |
15 |
|
100 ng |
14 |
14 |
|
500 ng |
12 |
13 |
|
1 μg |
11 |
12 |
6 Other Materials
6.1 DNA purification magnetic beads: Hieff NGS™ DNA Selection Beads (Yeasen Cat#12601) or AMPure® XP Beads (A63880) or other equivalent products.
6.2 RNA quality control: Agilent 2100 Bioanalyzer RNA 6000 Nano/Pico Chip or other equivalent products.
6.3 Adapters: Long adapters with Index (Yeasen Cat#12615-12618) or short adapter without Index (Yeasen Cat#12414-12415).
6.4 Library quality analysis: Agilent 2100 Bioanalyzer DNA 1000 Chip/ High Sensitivity Chip or other equivalent products; library quantitative reagents.
6.5 Other materials: absolute ethanol, sterile ultrapure water, low retention pipette tips, PCR tube, magnetic stands, thermal cycler, etc.
Workflow
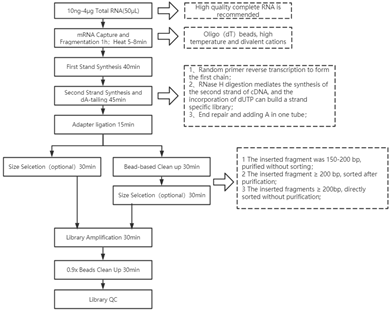
Figure 1. The workflow of mRNA library construction kit
Catalog No.:*
Name*
phone Number:*
Lot:*
Email*
Country:*
Company/Institute:*
Recommended products