The essence of NGS library quantification is nucleic acid (DNA or RNA) quantitation. Accurate analysis of NGS library quantification is crucial to obtain high-quality sequencing data. If the quantitative concentration of the NGS library is high, the yield of sequencing data may be low due to insufficient clustering. However, low quantitative concentration may cause signal interference between clusters to produce low-quality data, and even lead to sequencing failure. In addition, with the increase in the sequencing flux of sequencers, multiple NGS libraries are often combined for sequencing. According to the requirements of the size of sequencing data, NGS libraries with corresponding molar quantities are mixed based on the quantitative results to stabilize the output of the size of each sample data. Therefore, accurate NGS library quantification of libraries is crucial.
At present, there are two main methods of NGS library quantification, the ultraviolet absorption method, and the fluorescent dye method. NanoDrop is a DNA quantification instrument designed based on the principle of ultraviolet absorption; Qubit is a DNA quantification instrument developed according to the principle of fluorescent dye. The quantitative accuracy and cost of these two methods are different. The qubit method is mainly recommended here. So why do we recommend the qubit to you? What are the characteristics of Qubit products supplied by Yeasen? Please continue to read...
1. Why do we recommend Qubit to you?
2. What are the characteristics of Qubit products supplied by Yeasen?
3. How to choose Qubit for different samples?
1. Why do we recommend Qubit to you?
There is no doubt that Qubit is better than Nanodrop. So why is qubit better than nanodrop? What is the difference between qubit and nanodrop? First, as mentioned before, they differ in the principle of DNA quantification, NanoDrop is designed based on ultraviolet absorption. The principle of the ultraviolet absorption method is that there are purine and pyrimidine bases in nucleic acid components, and these bases have conjugate double bonds. There is strong light absorption at 250-280nm in the UV region, and the maximum absorption value is about 260nm. Therefore, the UV absorption of nucleic acid can be used for concentration quantification. NanoDrop is simple and quick to operate, but it cannot distinguish between DNA, RNA, degraded nucleic acids, free nucleotides, and other impurities. It is not suitable for obtaining accurate library quantitative data. But, Qubit quantitation is the best choice for accurate NGS library quantification, especially for DNA quantitation. So what are the features of qubit DNA quantification? Please view the following table:
| Specificity | Fluorescent dyes preferentially bind dsDNA |
| The length of DNA fragment | No specific restriction |
| Accuracy | High |
| Sample size requirements | Dilute samples of 1-20μl |
| The range of detection concentration | 0.2-100ng |
| The number of standards | Usually two |
| Experiment with manual operation time | Set the time to 5 minutes and the measurement time for a single sample to 3s |
| Detection time | 5min-30min (depending on sample size) |
| Applicable instrument | Qubit (3.0/4.0); Microplate reader with the corresponding wavelength |
| Applicable scene | dsDNA quantification; dsDNA quantification in NGS library construction and rapid quantification in conventional library |
2. What are the characteristics of Qubit products supplied by Yeasen?
Before introducing the characteristics of Qubit products supplied by Yeasen, let's understand how it works.
2.1 How does Qubit work?
The principle of Qubit is to use fluorescent dyes to bind to DNA molecules to emit fluorescence, and then measure the fluorescence intensity to obtain the concentration of nucleic acid. Fluorescent dyes fluoresce only after binding to the DNA double-strand, and the resulting fluorescence is proportional to the DNA concentration.
2.2 Popular product supplied by Yeasen
The 1×dsDNA HS Assay Kit for Qubit® (Cat#: 12642ES), developed on the basis of the dsDNA HS Assay Kit for Qubit® kit (Cat#: 12640ES), is a ready-to-use Qubit premix solution to predilute the concentrated dyes in the original kit into a working solution. When operating, you only need to mix the working liquid with the sample to be tested to determine the sample concentration by Qubit® fluorometer, which is more convenient to use.
2.3 Product Features
Simple and easy to operate: ready-to-use premixed liquid, no need to prepare working liquid, simple operation, quantitative more time saving;
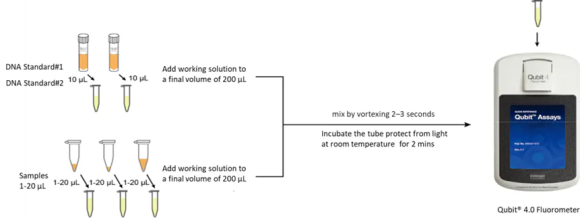
Figure 1. The processes of dsDNA HS Assay Kit for Qubit
Ultra-high sensitivity: 64x gradient dilution precise quantification, 0.2-100 ng dsDNA has good linear relationship;
Super specificity: Specific detection dsDNA, has excellent tolerance to some conventional pollutants;
Fast binding speed of dye: saturation can be reached within 2 min, so there is no need to wait for rapid reading of accurate quantitative results;
High stability: The fluorescence signal lasted for 3h, and the quantitative accuracy was not affected after 20 days at room temperature.
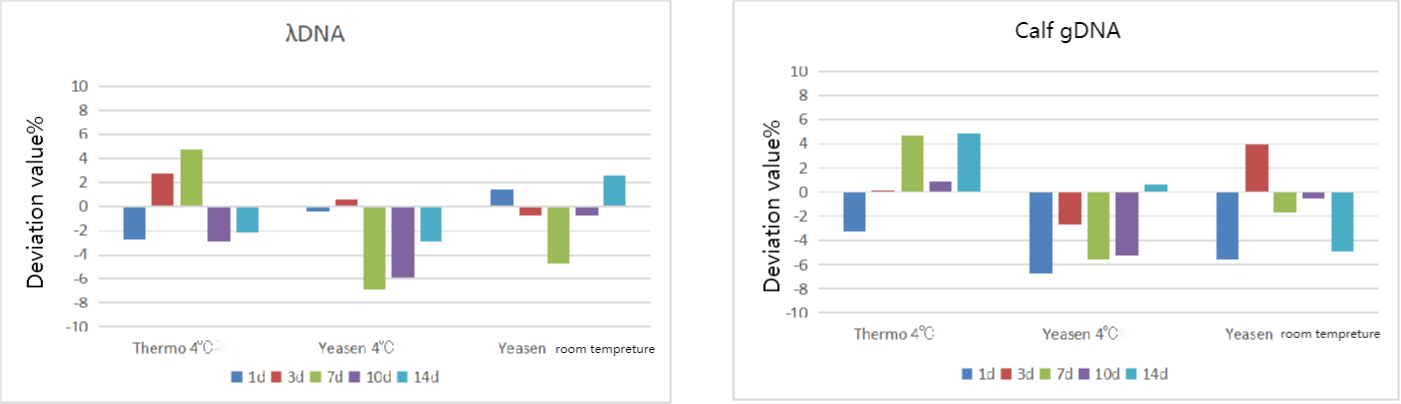
Figure 1. The product stability test. The results show it remains stable after being stored at room temperature for 2 weeks (the deviation between the measured value and the theoretical value was <10%)
Note: Two concentrations of dsDNA were selected, and Thermo#Q33230 stored at 4℃, Yeasen#12642 stored at 4℃, and Yeasen#12642 stored at room temperature (about 25℃) were used to determine the concentration values at different times, and the deviation rate between each measurement and the first one (denoted as 0d) was calculated.
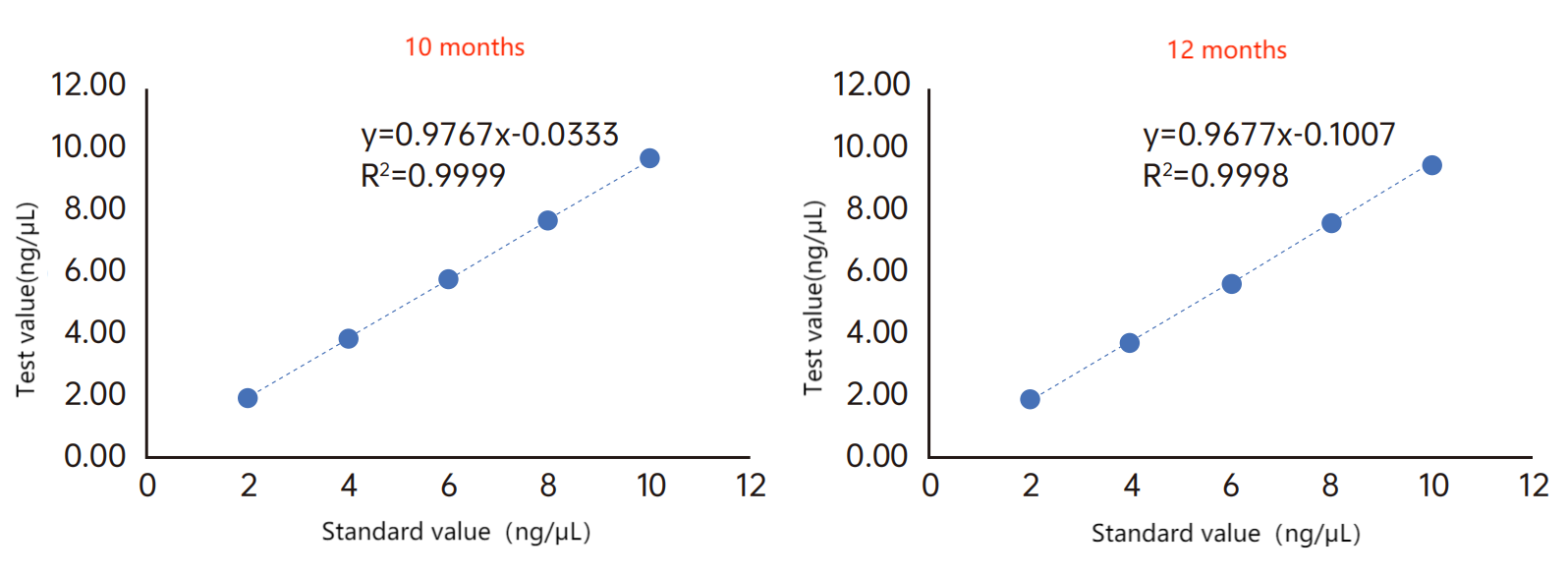
Figure 2. Product stability test. Super stability, high relevance of test results within the validity period
3. How to choose Qubit for different samples?
Qubit, fast quantification! Using Qubit for library quantification has many advantages. Its detection process is fast and effective. Sequencing results can be obtained within a few seconds for a single sample and the concentration can be directly output, which is very intuitive. At the same time, it can also be compatible with most of the enzyme label instrument for batch sample quantification, which is the first choice for conventional library quantification and dsDNA quantification.
| Product Name | Catalog No. | Size | Application |
| 1×dsDNA HS Assay Kit for Qubit® | 12642ES | 100 T/500T |
dsDNA quantification Library quantification |
| dsDNA HS Assay Kit for Qubit® | 12640ES | 100 T/500T | |
| ssDNA Assay Kit | 12645ES | 100 T/500T |
ssDNA quantification Library quantification |